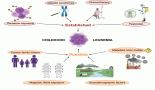
Introduction: T-lymphoblastic leukaemia accounts for approximately one-fourth of acute lymphoblastic leukaemia cases. Sequencing approaches have identified >100 genes that can be mutated in T-cell acute lymphoblastic leukaemia (T-ALL). However, the revised WHO 2022 edition of lymphoid neoplasms still does not incorporate molecular signatures into the T-ALL subgrouping unlike B-ALLs and acute myeloid leukemia, which are classified mainly based on molecular landscapes.
Methods: This retrospective observational study included all newly diagnosed patients of T-lymphoblastic leukaemia of all age groups who presented during the period between January 2022 and October 2023 in whom complete baseline diagnostic work-up was available including flow cytometry, fluorescence in situ hybridization and next generation sequencing studies.
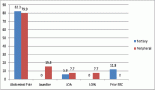
Results: There was a lower frequency of karyotypic abnormalities in adult early T progenitor (ETP)-ALLs than in other sub-groups. Non-ETP ALLs showed significant association with NOTCH1 mutations (p ≤ 0.00001), followed by JAK3 (p = 0.01), FBXW7 (p = 0.066) and PHF6 (p = 0.09) mutations. There was no difference between adult and pediatric patients, in terms of genomic profiling except in the PHF6 gene. There was no significant difference between NOTCH1-mutated and NOTCH1-wild T-ALL patients as well as NOTCH1-heterodimerization versus NOTCH1-PEST mutated patients in terms of measurable residual disease (MRD), relapse-free survival (RFS) and/or overall survival (OS). 45.1% of all TALL patients harboured ≥3 mutations. However, the complex molecular profile did not correlate significantly with MRD positivity and poor RFS and/or OS rates.
Conclusion: Molecular profiling of TALLs do not significantly impact long-term survival outcomes. In resource-constrained settings, we can get away by not doing comprehensive molecular profiling of TALLs at baseline and restrict the sequencing assay to only those cases that are persistently MRD positive or have relapsed.